Chinese Journal of Tissue Engineering Research ›› 2015, Vol. 19 ›› Issue (7): 1087-1093.doi: 10.3969/j.issn.2095-4344.2015.07.019
Previous Articles Next Articles
Muscle-specific microRNA-206: research status and prospects
Zheng Pan-pan, Yu Xin-kai, Zuo Qun, Li Wan-wan, Song Ya-yun
- Shanghai University of Sport, Shanghai 200438, China
-
Online:2015-02-12Published:2015-02-12 -
Contact:Yu Xin-kai, M.D., Associate professor, Shanghai University of Sport, Shanghai 200438, China -
About author:Zheng Pan-pan, Studying for master’s degree, Shanghai University of Sport, Shanghai 200438, China -
Supported by:the National Natural Science Foundation of China, No. 31171139; the Graduate’s Education Innovation Fund of Shanghai University of Sport, No. yjscx2014038; a grant from Shanghai Key Laboratory for Development and Protection of Human Exercise Capacity (Shanghai University of Sport), Ministry of Education, No. 11DZ2261100
CLC Number:
Cite this article
Zheng Pan-pan, Yu Xin-kai, Zuo Qun, Li Wan-wan, Song Ya-yun . Muscle-specific microRNA-206: research status and prospects[J]. Chinese Journal of Tissue Engineering Research, 2015, 19(7): 1087-1093.
share this article
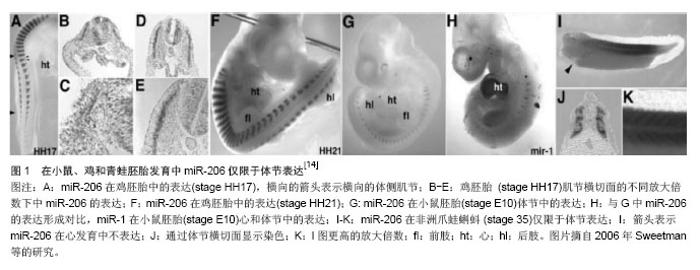
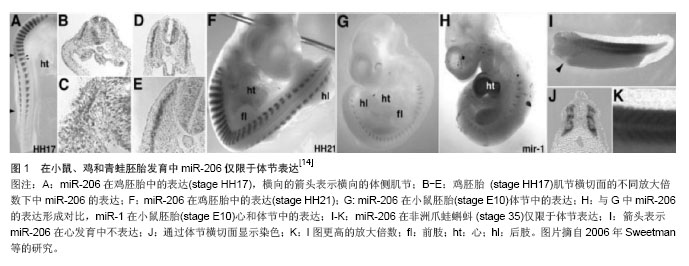
2.1 MicroRNA-206的生物合成和作用机制 2.1.1 MicroRNAs的生物合成 miRNA是一类非编码、内源性的小RNA,长21-25个核苷酸,主要调节基因转录后的表达[1],最初是1993年在线虫中发现——lin-4[2-3]。miRNA具有高度保守性,有研究预测多种miRNA都能在其他种系中找到同源体。在206种脊椎动物的研究中,已经发现了30 000多个成熟miRNA[4]。大量研究表明,骨骼肌的生成发展几乎都需要miRNA参与调控,特别是肌原性的miRNA。 目前,在动物体内miRNA的生物合成过程已经得到了较为详尽的诠释。MiRNA基因主要在RNA聚合酶Ⅱ的作用下转录成较长的初级转录产物pri-miRNA[5],有时 RNA聚合酶Ⅲ也会参与。Pri-miRNA在细胞内RNaseⅢ内切酶(Drose)的作用下加工合成70个碱基的pre-miRNA,pre-miRNA在细胞转运蛋白5(exportin 5)作用下被运输到细胞质。在细胞质中,pre-RNA在RNaseⅢ酶(Dicer)的作用下形成22个核苷酸大小的成熟miRNA。 2.1.2 MicroRNA-206的作用方式和调节机制 miRNA通过与靶基因3’UTR区互补配对,指导miRNA复合体对靶mRNA进行切割或者翻译抑制。miRNA抑制mRNA的翻译机制至今为止还不是很清楚,但能够确定的是,miRNA 5’端2-8个核苷酸对于miRNA与3’UTR的配对非常重要,因此把5’端的这个部位称之为“种子”序列[6-7]。miRNA抑制mRNA的翻译机制至今为止是个迷。目前认为,miRNA对靶基因的作用机制主要有3种:miRNA与靶mRNA 完全互补结合,切割靶mRNA;miRNA与靶mRNA不完全互补结合,抑制靶蛋白翻译而不影响mRNA的稳定性;同时具有以上两种作用模式。动物的miRNA与靶序列的配对性较低,多数是对mRNA进行翻译抑制。 很多miRNA在某些特定的组织里高表达,在骨骼肌内特异性表达的miRNA的叫骨骼肌特异性miRNA,包括miR-1、miR-133a、miR -133b、miR -206、miR-208、miR-208b、miR-486和miR-499等[8-12]。这个亚类miRNA在肌肉组织中特异存在,参与肌肉形成过程中细胞的增殖和分化。MiR-206是骨骼肌特异性miRNA,是2003年从人和小鼠肌肉组织中克隆鉴定出来的,人、小鼠和大鼠的miR-206分别位于第6、1和9号染色体上[13],长度为22个核苷酸,是唯一只在骨骼肌中特异性表达的miRNA[14],在小鼠、鸡和青蛙胚胎发育中miR-206仅限于体节表达[14-15](见图1)。2006年chen等[16]通过基因芯片检测到miR-206在骨骼肌细胞分化成熟过程中表达增加且呈现一定的时空特异性。"

目前已发现多种转录因子能够激活并促进miRNA-206的转录过程。MRFS(生肌调节因子)-比较典型的是MyoD和bHLH,它们能通过调节miRNA-206来控制肌肉发育[17-18]。肌细胞分化因子MyoD能够促进横纹肌特异性miRNA的表达,刺激miR-206的表达能激活骨骼肌分化。Kim等[19]发现miR-206可通过下调MyoD的抑制因子MyoR等上调MyoD基因表达。转化生长因子β能通过抑制miR-206来改变HDAC4的表达[20]。 一个miRNA可以被多个靶基因所调控,同时一个miRNA可以调节多个靶基因。缝隙链接蛋白Cx43是miR-1/miR-206的靶基因[5,19],而Cx43的下调是成肌细胞融合的必要条件[21]。受体酪氨酸激酶(c-Met)是miR-206的靶基因,在横纹肌肉瘤细胞株中被确定,c-Met过度表达的肿瘤中,细胞异常地增殖和迁移,而miR-206能作为一种强有力的肿瘤抑制剂[22-23]。MiR-206另一个靶基因是HDAC4[16,24],是肌肉基因转录表达的抑制位点,HDAC4能部分介导miR-206的功能,在人体神经损伤后能促进神经突触的恢复,延缓肌萎缩侧索硬化症模型老鼠发病的进程[24]。miR-1和miR-206的靶基因Pax7的主要作用是抑制卫星细胞增殖[25-27]。在C2C12成肌细胞系中,miR-206通过DNA聚合酶α来影响mRNA的表达,miR-206的下调会通过抑制DNA的合成从而影响肌细胞的分化。B-ind1和Mnd也是miR-206的靶基因[19]。MiR-206可以通过调节基因-Hmgb3表达和DNA聚合酶的合成,抑制肌细胞的增殖、促进分化[28]。更多的miR-206的靶基因正在被预测和验证,包括卵泡抑素样1(Fstl1)[29],抗肌萎缩相关蛋白(Utrn)[29],和组织型基质金属蛋白酶抑制因子3(TIMP3)[30]。鉴定miRNA的靶基因是研究miRNAs功能发挥的关键,但现在仍是一个技术挑战。 2.2 MicroRNA-206在疾病中的表达 2.2.1 MicroRNA-206在骨骼肌疾病中的表达 研究证实,多种miRNA在心肌和骨骼肌疾病中表达失调,个别miRNA的表达异常可引起或减缓疾病的发生和进展。如在小鼠不同的发展阶段和不同组织进行Dicer的敲除,在胚胎早期Dicer酶缺陷的小鼠,肌肉质量的减少,肌细胞凋亡的增加,伴随着非正常的肌纤维,证明了miRNA在肌肉发育中扮演的关键性角色[31-33]。 杜氏肌营养不良(DMD)的患者体内,miR-223、miR-449和miR-206表达上升。Dystrophin基因缺失的mdx小鼠是研究肌营养不良发病机制和骨骼肌损伤再生机理的常用模型[34-35]。Mccarthy等[9]通过原位杂交显示,miR-206在mdx小鼠的膈膜和再生形成的肌肉中表达上升,而在完整的肌纤维中没有出现,暗示miR-206可能参与了肌肉损伤后的修复过程,提示了miR-206可能在促进肌纤维再生中有重要作用。Liu等[36]研究发现,4周龄mdx老鼠和正常老鼠相比,骨骼肌发生坏死和变性,mdx老鼠体内miR-206水平显著上升,并集中在新生肌管和再生的肌纤维中,而完整的肌纤维中并没有出现[27,37]。这些研究都显示miR-206在骨骼肌损伤再生的病理过程中发挥着重要作用。 强直性肌营养不良1型(DM1)的大部分患者,miR-206表达显著上升。但有趣的是,DM1患者和正常对照组相比,与miRNA-206有关的靶基因、mRNA和抗肌萎缩相关蛋白的水平都没有显著性差异[38]。Liu等[39]对肌营养不良的受试者进行干细胞疗法时发现,低氧水平能激活Notch信号通路,通过Hes/Hey蛋白来抑制miR-206和miR-1的表达,从而增加Pax7的表达,提高干细胞的自我更新和存活能力。 横纹肌肉瘤是一种恶性软组织瘤,由骨骼肌细胞分化程度低导致,现有治疗横纹肌肉瘤的方法具有明显的毒性,寻找一种新的治疗方法来治疗横纹肌肉瘤十分重要。MiR-206的水平和横纹肌肉瘤的临床表现有一定的相关性,横纹肌肉瘤患者骨骼肌内miR-206水平明显比正常人低[13]。MacQuarrie等[40]证实,miR-206能调控细胞从生长向分化转化。同时,Missiaglia等[41]发现miR-206的低表达会激活MAPK和核因子kB信号通路,而miR-206的超表达能通过转变横纹肌肉瘤细胞中总mRNA的表达来促进肌肉的分化,抑制细胞的增殖和迁移。 Williams等[24]证明在神经退行性疾病--肌萎缩侧索硬化症大鼠模型体内miR-206的表达上升,并且发现缺乏miR-206会加重肌萎缩侧索硬化症老鼠的病情,促进神经肌肉突触的变性。MiR-206对肌萎缩侧索硬化症的保护性效应可通过作用于靶位点HDAC4实现,这反过来又刺激FGF信号通路,促进损伤的运动神经元神经肌肉突触代偿性增生,从而减缓肌萎缩侧索硬化症进程[24]。也有研究显示,miR-206在神经肌肉再生过程中表达上升,通过作用于不同的分子,包括脑源性神经营养因子和它的受体(P75NTR)[42],促进肌纤维和运动神经元轴突之间的交流,从而减缓肌萎缩侧索硬化症的病情发展。最近,Valdez等[43]也证明,miR-206能调节神经肌肉损伤后神经肌肉接点的恢复。MiR-206对肌萎缩侧索硬化症的修复作用提示人们,控制miR-206的水平可能成为治疗肌萎缩侧索硬化症的新方法。 2.2.2 MicroRNA-206在其他疾病中的表达 人类很多疾病都与miRNA有密切联系[44]。哺乳类动物的miRNA主要是通过减少目标mRNA的翻译水平来抑制基因的表达,估计超过60%的人类蛋白质的基因编码都要在miRNA的调节下进行[45]。 生理条件下miR-206是骨骼肌特异性miRNA[14],如在正常人大脑内miR-206的表达在一个比较低的水平[46],在健康的心脏中miR-206的水平也几乎检测不到[47]。而在患某些疾病情况下,miR-206的水平会增加,如在心肌梗死、糖尿病型心肌病或者其他心脏疾病的情况下[47]。MiR-206已被认为一种抑癌基因,在肾细胞癌和子宫内腺癌中均发现miR-206表达上调能抑制细胞的增殖和转移[48-49]。 2.3 MicroRNA-206在骨骼肌损伤和运动中的研究 2.3.1 MicroRNA-206在肌肉损伤中的研究 骨骼肌损伤能激活成熟的肌原性干细胞-卫星细胞来进行骨骼肌的再生。骨骼肌损伤早期,卫星细胞中miR-206下调,这种现象是因为早期阶段miR-1和miR-206受到卫星细胞的抗增殖作用[50],pax7水平上升增加卫星细胞的增殖能力,损伤后期,miR-206水平上升,促进卫星细胞的分化和肌管的融合,揭示了miR-206在骨骼肌再生中的重要作用。MiR-206在骨骼肌损伤中的研究(表1)。 "


Mccarthy等[8]通过切除协同肌造成跖肌肥大,作者发现实验组miR-206和pri-206的表达不同步,pri-miR-206是对照组的18.3倍,miR-206表达水平没有显著性改变。提示miR-206可能与骨骼肌功能性肥大有关。因此,后续研究可以检测骨骼肌肥大后期时间点miR-206的表达是否会上升。 众多研究结果显示,骨骼肌严重损伤后miR-206的变化比较明显。Nakasa等[52]用手术刀切割损伤部分大鼠胫骨前肌肌腹,发现大鼠胫骨前肌撕裂7 d内,miR-206表达先下降再上升。他们还发现,给对照组注射siRNA,实验组局部注射相同剂量双链miR-1,miR-133和miR-206的混合物。在注射后的3-7d,发现注射miR-1,miR-133和miR-206的混合物能促进MyoD1、myogenin和 Pax7 mRNA和蛋白质的表达。1周后发现实验肌肉内形态学和生理学功能的再生,纤维化程度也比对照组小。说明外源性的注射miR-1、miR-133和miR-206能诱导肌原性标记物的表达,加速大鼠骨骼肌的再生。同时,在体外培养中发现,加入miR-1,miR-133和miR-206的混合物,能促进肌管的分化,同时MyoD1、myogenin和Pax7的表达上升。这些实验都证明了miR-1,miR-133和miR-206在骨骼肌生长和发育过程中起着至关重要的调节作用。Liu等[36]也发现miR-206的缺乏,会导致卫星细胞延迟分化,miR-206的缺乏会加剧mdx小鼠营养不良的状况,延迟肌肉的再生和成熟。因此,局部注射miR-1,miR-133和miR-206可能成为治疗骨骼肌损伤的新方法。 Roberts等[53]在正常小鼠胫骨前肌注射25 µL 10-5 mol/L的心脏毒素,15 min、4 d、5 d后,虽然小鼠血清中miR-206的水平表达没有显著性差异,但在注射心脏毒素后15 min miR-206的表达有一定的上升趋势。作者认为创伤后的血清取样密度不够频繁,并不能真正代表创伤后血清中miR-206的表达变化状况。研究还发现,胫骨前肌内miR-206及其转录因子的表达和血清中有一定的相关性,暗示了细胞外的miRNA可能成为评测骨骼肌再生状态的指标。 Yuasa等[37]在mdx小鼠胫骨前肌注射CTX造成损伤后,miR-206水平先下降再升高,第5天miR-206的水平是损伤前的10倍。MiR-1和MiR-133a在损伤后先下降再上升,到第4周恢复到注射前水平。同时Yuasa还通过原位杂交发现,miR-206主要集中在新生的肌管、再生的肌肉和不成熟的肌纤维中,而未受损的肌纤维和小单核细胞中没有miR-206的表达,证实了miR-206与肌肉的再生有关。 Liu等[36]做了类似研究,对正常小鼠胫骨前肌注射CTX发现,miR-206的表达也是呈先下降再升高再下降趋势,有趣的是,正常小鼠在30 d时miR-206恢复到了实验前水平,而Yuasa等的mdx小鼠在损伤8周后仍保持在一个较高的水平,进一步支持了miR-206能促进肌肉的再生。 2.3.2 MicroRNA-206在运动中的研究 在体育训练领域,需要新的生物学标记来评价运动量和运动负荷等情况。近年来,众多研究探索了短期和长期运动对miR-206在肌肉和血液中的影响,暗示了miR-206可能成为运动训练、运动损伤等方面的指标。MiR-206在运动中的研究(表2)。 Drummond等[54]对没有训练的男性进行研究,发现青年与老年受试者进行一次抗阻运动3 h和6 h后,青年和老年受试者骨骼肌中pri-miR-206的表达均增加,而miR-206运动前后表达无显著差异。MiR-1的表达水平在青年和老年受试者的骨骼肌中均升高,而miR-133a和miR-206的表达水平均没有变化,其具体机制仍待研究。 长期耐力运动会引起体内miR-206表达的变化。Nielsen等[55]对10名健康男青年进行12周的耐力训练后发现,股外侧肌miR-1、miR-133a、miR-133b和miR-206显著下降,特别是miR-206,减少了49%。这些miRNA的改变在停训后2周回到训练前的水平,而miR-206是如何调节骨骼肌适应耐力运动的机制还不是很清楚。有趣的是,在进行耐力训练前后,分别对受试者进行60 min的65%最大功率的自行车运动,发现运动后即刻、1 h和3 h,miR-206都表达没有显著性差异。这些数据说明miR-206在中等强度生理性刺激之后不会产生变化,但实验不能排除在之后的时间点可能出现的miR-206的变化。 同时,Russell等[56]也做过类似的实验,受试者为9名没有受过训练的健康男子,进行3h的中等强度耐力运动之后,miR-1、miR-133a、miR-133b表达都有上升,而miR-206表达没有变化。接着Russel让受试者经过 10 d的耐力训练后,进行肌肉活检,发现miR-1和miR-31的表达增加,miR-206的表达没有显著性差异。这项实验与Nielsen等[50]的研究相比,不同的地方是运动量和进行长期锻炼持续的时间,这项研究的受试运动量更小,而且锻炼的持续时间比较短,这可能是导致长期运动后两组受试者肌肉miR-206不同变化趋势的原因。 运动后miR-206在血液中的研究在近几年才开始。2008年,有研究首次探索了血液中miRs的改变及其分布情况。之后对血液中miRs水平的变化研究越来越多。Kroh等[60]研究分析了在血清和血浆中miRs分布的差异。Gomes等[58]检测了受试者血液中3种miRNA的变化,发现在半程马拉松后,受试者血浆中的miR-1、miR-133a和miR-206表达上升。Mooren等[59]做了类似的实验,检测了14名男性耐力运动员在马拉松赛后血液中miRs表达的变化。发现血液中miR-499、miR-208、miR-1、miR-133a、miR-206水平在运动后即刻显著性上升,miR-206水平上升幅度最大,运动后即刻大概是运动前的20倍,并且miR-206在运动后24 h变化幅度虽然有所下降,但没有恢复到运动前水平。Aoi等[57]对11名年轻男性进行进行60min70%VO2max的一次性运动,运动后进行测量血清中miR-1、miR-133a、miR-133b、miR-206、miR-208b、miR-486和miR-499的检测,发现只有miR-486一次性运动和长期运动前后有显著性变化,miR-206没有显著性变化。 结合上述研究发现,miR-206在肌肉和血液中的水平对一次性强度不是很大的运动不敏感,但在马拉松比赛这种负荷比较大的运动后水平会显著性上升。miR-206长期运动适应后水平下降,但当运动适应停训一段时间后,miR-206会恢复到原先的水平,而其机制还有待研究。 "

| [1]Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116(2):281-297. [2]Lee RC, Feinbaum RL, Ambros V,et al. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell.1993;75(5):843-854. [3]Wightman B, Ha I, Ruvkun G. Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell. 1993;75(5):855-862. [4]Kozomara A, Griffiths-Jones S. miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res.2011;39(Database issue): D152-D157. [5]Kim VN, Han J, Siomi MC. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol. 2009;10(2):126-139. [6]Fabian MR, Sonenberg N, Filipowicz W.Regulation of mRNA translation and stability by microRNAs. Annu Rev Biochem. 2010;79:351-379. [7]Thomas M, Lieberman J, Lal A. Desperately seeking microRNA targets. Nat Struct Mol Biol.2010;17(10): 1169-1174. [8]Mccarthy JJ, Esser KA. MicroRNA-1 and microRNA-133a expression are decreased during skeletal muscle hypertrophy. J Appl Physiol (1985). 2007;102(1):306-313. [9]Mccarthy JJ, Esser KA, Andrade FH.MicroRNA-206 is overexpressed in the diaphragm but not the hindlimb muscle of mdx mouse. Am J Physiol Cell Physiol. 2007;293(1): C451-C457. [10]Small E M, O'Rourke J R, Moresi V, et al. Regulation of PI3-kinase/Akt signaling by muscle-enriched microRNA-486. Proc Natl Acad Sci U S A. 2010;107(9):4218-4223. [11]van Rooij E, Liu N, Olson EN. MicroRNAs flex their muscles. Trends Genet. 2008; 24(4):159-166. [12]Callis TE, Deng Z, Chen JF, et al. Muscling through the microRNA world. Exp Biol Med (Maywood). 2008;233(2): 131-138. [13]Cifuentes D, Xue H,Taylor DW,et al.A novel miRNA processing pathway independent of Dicer requires Argonaute2 catalytic activity. Science.2010;328(5986): 1694-1698. [14]Mccarthy JJ. MicroRNA-206: the skeletal muscle-specific myomiR. Biochim Biophys Acta.2008;1779(11):682-691. [15]Sweetman D, Rathjen T, Jefferson M, et al. FGF-4 signaling is involved in mir-206 expression in developing somites of chicken embryos. Dev Dyn.2006;235(8):2185-2191. [16]Chen JF, Mandel EM, Thomson JM, et al. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation.Nat Genet.2006;38(2): 228-233. [17]Sweetman D, Goljanek K, Rathjen T, et al.Specific requirements of MRFs for the expression of muscle specific microRNAs, miR-1, miR-206 and miR-133. Dev Biol. 2008; 321(2):491-499. [18]Rao PK, Kumar RM, Farkhondeh M, et al. Myogenic factors that regulate expression of muscle-specific microRNAs. Proc Natl Acad Sci U S A.2006;103(23):8721-8726. [19]Kim H K, Lee Y S, Sivaprasad U, et al. Muscle-specific microRNA miR-206 promotes muscle differentiation. J Cell Biol.2006;174(5):677-687. [20]Winbanks CE, Wang B,Beyer C, et al.TGF-beta regulates miR-206 and miR-29 to control myogenic differentiation through regulation of HDAC4. J Biol Chem.2011; 286(16): 13805-13814. [21]Anderson C, Catoe H, Werner R. MIR-206 regulates connexin43 expression during skeletal muscle development. Nucleic Acids Res.2006;34(20):5863-5871. [22]Yan D, Dong XE, Chen X,et al.MicroRNA-1/206 targets c-Met and inhibits rhabdomyosarcoma development. J Biol Chem. 2009;284(43):29596-29604. [23]Taulli R, Bersani F, Foglizzo V, et al. The muscle-specific microRNA miR-206 blocks human rhabdomyosarcoma growth in xenotransplanted mice by promoting myogenic differentiation. J Clin Invest.2009;119(8):2366-2378. [24]Williams AH, Valdez G, Moresi V, et al.MicroRNA-206 delays ALS progression and promotes regeneration of neuromuscular synapses in mice. Science.2009; 326(5959): 1549-1554. [25]Tang F, Kaneda M, O'Carroll D, et al.Maternal microRNAs are essential for mouse zygotic development. Genes Dev.2007; 21(6): 644-648. [26]Dey BK, Gagan J, Dutta A. miR-206 and-486 induce myoblast differentiation by downregulating Pax7. Mol Cell Biol. 2011; 31(1):203-214. [27]Cacchiarelli D, Martone J, Girardi E, et al. MicroRNAs involved in molecular circuitries relevant for the Duchenne muscular dystrophy pathogenesis are controlled by the dystrophin/nNOS pathway. Cell Metab.2010;12(4):341-351. [28]Maciotta S, Meregalli M, Cassinelli L, et al. Hmgb3 is regulated by microRNA-206 during muscle regeneration. PLoS One.2012;7(8):e43464. [29]Rosenberg MI, Georges SA, Asawachaicharn A, et al. MyoD inhibits Fstl1 and Utrn expression by inducing transcription of miR-206. J Cell Biol.2006;175(1):77-85. [30]Liu H,Chen SE,Jin B,et al.TIMP3: a physiological regulator of adult myogenesis. J Cell Sci.2010;123(Pt 17):2914-2921. [31]Bernstein E, Kim SY, Carmell MA, et al.Dicer is essential for mouse development. Nat Genet. 2003;35(3):215-217. [32]Kanellopoulou C, Muljo SA, Kung AL, et al. Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev.2005;19(4):489-501. [33]Murchison EP, Stein P, Xuan Z, et al. Critical roles for Dicer in the female germline. Genes Dev.2007;21(6):682-693. [34]Eisenberg I,Eran A,Nishino I,et al.Distinctive patterns of microRNA expression in primary muscular disorders. Proc Natl Acad Sci U S A. 2007;104(43):17016-17021. [35]Greco S, De Simone M, Colussi C, et al. Common micro-RNA signature in skeletal muscle damage and regeneration induced by Duchenne muscular dystrophy and acute ischemia. FASEB J. 2009;23(10):3335-3346. [36]Liu N, Williams AH, Maxeiner JM, et al. microRNA-206 promotes skeletal muscle regeneration and delays progression of Duchenne muscular dystrophy in mice. J Clin Invest.2012;122(6):2054-2065. [37]Yuasa K, Hagiwara Y, Ando M, et al. MicroRNA-206 is highly expressed in newly formed muscle fibers: implications regarding potential for muscle regeneration and maturation in muscular dystrophy.Cell Struct Funct. 2008;33(2):163-169. [38]Gambardella S, Rinaldi F, Lepore SM, et al. Overexpression of microRNA-206 in the skeletal muscle from myotonic dystrophy type 1 patients. J Transl Med. 2010;8:48. [39]Liu W, Wen Y, Bi P, et al. Hypoxia promotes satellite cell self-renewal and enhances the efficiency of myoblast transplantation. Development. 2012;139(16):2857-2865. [40]Macquarrie KL, Yao Z, Young JM, et al. miR-206 integrates multiple components of differentiation pathways to control the transition from growth to differentiation in rhabdomyosarcoma cells. Skelet Muscle.2012;2(1):7. [41]Missiaglia E, Shepherd CJ, Patel S, et al. MicroRNA-206 expression levels correlate with clinical behaviour of rhabdomyosarcomas. Br J Cancer. 2010;102(12):1769-1777. [42]Miura P,Amirouche A,Clow C,et al. Brain-derived neurotrophic factor expression is repressed during myogenic differentiation by miR-206. J Neurochem. 2012;120(2):230-238. [43]Valdez G, Heyer MP, Feng G, et al.The role of muscle microRNAs in repairing the neuromuscular junction. PLoS One. 2014;9(3):e93140. [44]Novak J, Kruzliak P, Bienertova-Vasku J, et al.MicroRNA-206: a promising theranostic marker. Theranostics. 2014;4(2): 119-133. [45]Guo H, Ingolia NT, Weissman JS, et al. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature.2010;466(7308):835-840. [46]Lehotzky A, Lau P, Tokesi N, et al.Tubulin polymerization- promoting protein (TPPP/p25) is critical for oligodendrocyte differentiation. Glia.2010;58(2):157-168. [47]Westendorp B, Major JL, Nader M, et al.The E2F6 repressor activates gene expression in myocardium resulting in dilated cardiomyopathy. FASEB J. 2012;26(6):2569-2579. [48]Hidaka H, Seki N, Yoshino H, et al. Tumor suppressive microRNA-1285 regulates novel molecular targets: aberrant expression and functional significance in renal cell carcinoma. Oncotarget. 2012;3(1):44-57. [49]Chen X,Yan Q,Li S,et al. Expression of the tumor suppressor miR-206 is associated with cellular proliferative inhibition and impairs invasion in ERalpha-positive endometrioid adenocarcinoma. Cancer Lett. 2012;314(1):41-53. [50]Chen J F, Tao Y, Li J, et al. microRNA-1 and microRNA-206 regulate skeletal muscle satellite cell proliferation and differentiation by repressing Pax7. J Cell Biol.2010;190(5): 867-879. [51]Mccarthy JJ, Esser KA, Peterson CA, et al. Evidence of MyomiR network regulation of beta-myosin heavy chain gene expression during skeletal muscle atrophy. Physiol Genomics. 2009;39(3):219-226. [52]Nakasa T,Ishikawa M, Shi M, et al. Acceleration of muscle regeneration by local injection of muscle-specific microRNAs in rat skeletal muscle injury model. J Cell Mol Med. 2010; 14(10):2495-2505. [53]Roberts TC, Godfrey C, Mcclorey G, et al. Extracellular microRNAs are dynamic non-vesicular biomarkers of muscle turnover. Nucleic Acids Res.2013;41(20):9500-9513. [54]Drummond MJ, Mccarthy JJ, Fry CS, et al. Aging differentially affects human skeletal muscle microRNA expression at rest and after an anabolic stimulus of resistance exercise and essential amino acids. Am J Physiol Endocrinol Metab. 2008; 295(6):E1333-E1340. [55]Nielsen S, Scheele C, Yfanti C, et al.Muscle specific microRNAs are regulated by endurance exercise in human skeletal muscle.J Physiol.2010;588(Pt 20):4029-4037. [56]Russell AP, Lamon S, Boon H, et al.Regulation of miRNAs in human skeletal muscle following acute endurance exercise and short-term endurance training. J Physiol. 2013;591(Pt 18): 4637-4653. [57]Aoi W, Ichikawa H, Mune K, et al. Muscle-enriched microRNA miR-486 decreases in circulation in response to exercise in young men. Front Physiol.2013;4:80. [58]Gomes C P, Oliveira-Jr G P, Madrid B, et al. Circulating miR-1, miR-133a, and miR-206 levels are increased after a half-marathon run. Biomarkers. 2014;19(7):585-589. [59]Mooren FC, Viereck J, Kruger K,et al.Circulating microRNAs as potential biomarkers of aerobic exercise capacity. Am J Physiol Heart Circ Physiol.2014; 306(4):H557-H563. [60]Kroh EM, Parkin RK, Mitchell PS, et al.Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods.2010;50(4): 298-301. |
| [1] | Zhang Shuang, Tan Rui, Wang Chunxiao, Wu Fengyu, Guo Hongyu. MicroRNAs for assessing the motion control of human skeletal muscles [J]. Chinese Journal of Tissue Engineering Research, 2021, 25(17): 2755-2760. |
| [2] | Ren Chunmei, Liu Yufang, Xu Nuo, Shao Miaomiao, He Jianya, Li Xiaojie. Non-coding RNAs in human dental pulp stem cells: regulations and mechanisms [J]. Chinese Journal of Tissue Engineering Research, 2020, 24(7): 1130-1137. |
| [3] | Han Mingli, Lü Pengwei, Qian Xueke, Yang Xue, Yang Yunqing, Gu Yuanting. MicroRNA-10b regulates aldehyde dehydrogenase 1 mRNA and protein expression in breast cancer MCF-7 cell line [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(9): 1349-1353. |
| [4] | Zhang Yuantong. Body composition and bone mineral density of juvenile basketball players versus ordinary middle school students: data from a middle school [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(3): 341-347. |
| [5] | Lan Qingshi. Participation degree and control ability of human latissimus dorsi, trapezius, triceps, musculus and deltoid during cross support of hand ring: analysis on contribution rate of major muscle group [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(3): 361-366. |
| [6] | Gao Hongqiang, Liu Jing, Li Zhiqiang, Wang Hailei, Zhao Xiongqi, Zhang Shengning, Ran Jianghua, Li Li . Ulinastatin improves rat liver metabolism after reduced-size liver transplantation [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(3): 435-440. |
| [7] | Kong Lingyao, Li Tao, Zeng Xinglin, Li Jian, Xiong Yan. Synovial chondromatosis: how to improve the diagnosis accuracy and clearance rate of tumor cells [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(28): 4570-4575. |
| [8] | Yu Hong, Liu Yan. Strength of knee flexor and extensor in football athletes after vibration training [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(15): 2327-2331. |
| [9] | Zhu Yaojia1, Huo Hongfeng1, 2. Balance ability and foot type characteristics during different postures of standing [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(15): 2345-2349. |
| [10] | Deng Guiying1, 2, Zeng Gaofeng3, Cen Zhongxi1, Gao Yunbing1, Cao Baichuan1, Huang Jianhua1, Zong Shaohui1 . Effect of miRNA-136-5p on inflammatory factors in rat models of acute spinal cord injury [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(15): 2397-2402. |
| [11] | Li Xintong1, Pan Weimin2, Qin Huasheng1, Qu Lei1, Zhang Hengyin1, Zhu Xinrui1. Blood flow restriction training: a new method for accelerating musculoskeletal rehabilitation [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(15): 2415-2420. |
| [12] | Wang Minjia1, Qi Ziyi2, Zhu Weihua1, Sun Junzhi1. Mechanism and clinical research advance of muscle atrophy: thinking based on the 65th Annual Meeting of American College of Sports Medicine [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(15): 2421-2426. |
| [13] | Gong Hewei, Liu Chengwei, Liang Bingsheng, Li Gang. Lentiviral vector-mediated overexpression of miRNA-1 in L6 myoblasts: cell proliferation, differentiation and histone deacetylase expression [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(13): 2075-2080. |
| [14] | Qi Minjun, Wu Xiaopeng, Zhou Zhongxing, Jiang Xiaodong. miR-142-3p effect on the stemness of bladder cancer stem cells via regulation of S1PR3 [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(13): 2028-2034. |
| [15] | Yuan Yiwen, Gui Zhu, Zhang Hongchao. Changes in muscle injury and pulse wave velocity after exercise with compression leg cuffs in college students without fitness habits [J]. Chinese Journal of Tissue Engineering Research, 2019, 23(11): 1687-1692. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||